Aim:
Reduce the dimension of a multivariate data set composed of a series of images.
The series of images may be composed of the different components of a multivariate image (the 3 components of a color image, for instance) or of a set of related images (a time series, for instance).
The series of images is reduced to a smaller series, where the different images of the new series are the principal components (or the factorial images) of the original series.
Method:
Dimensionality reduction is performed according to well-known algorithms (Principal Components Analysis or Correspondence Analysis).
For the moment, only PCA without data centering nor data normalization is implemented. Other variants will be made available soon.
User interface:
Input: The image series to be processed must be within a stack.
An example with four images (four "maps" of an alloy, representing the composition in carbon, chromium, nickel and titanium):
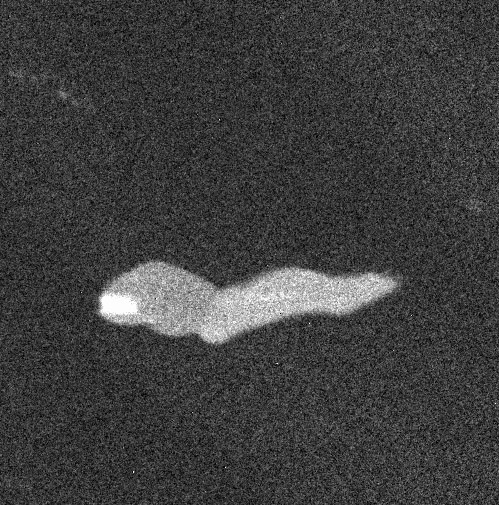
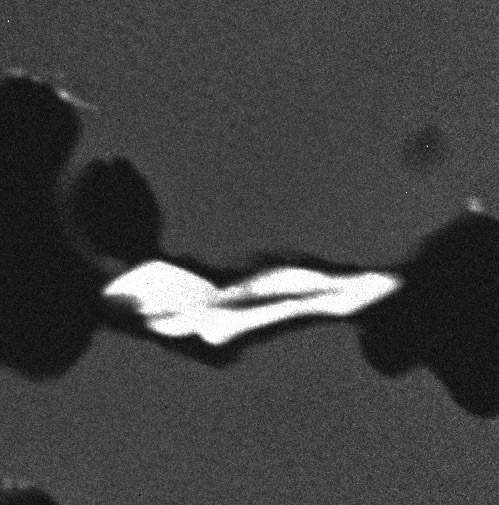
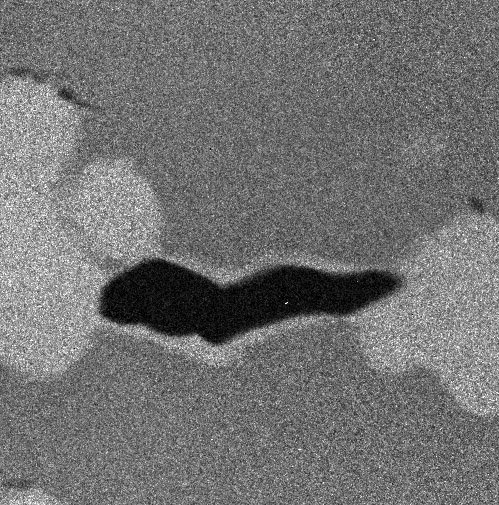
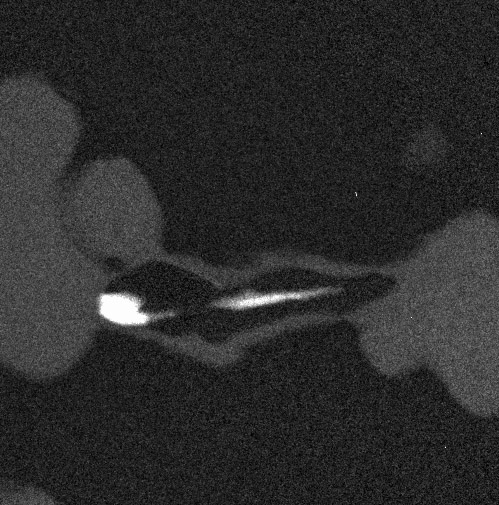
The 4 images are gathered in a stack:
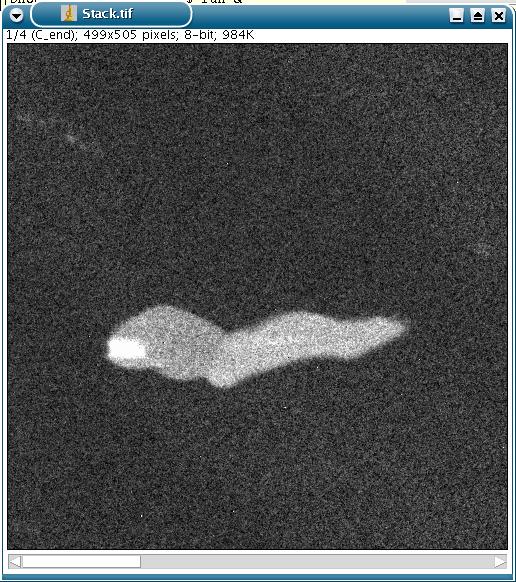
When the plug-in is activated, the user is prompted to select:
a) the method he/she wants to implement (at this moment, only PCA without centering nor mormalization, and CA, are implemented!!!)
b) the number of factorial images he/she wants to get as an output.
 or
or 
The resultant factorial images are put within a stack.
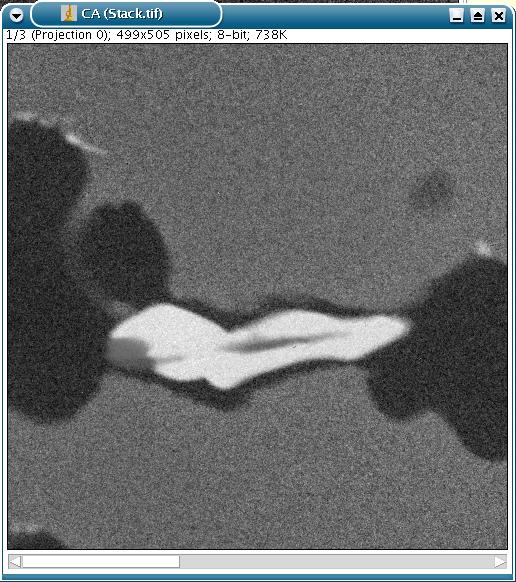
Of course, the stack can be split into factorial images if necessary.
The scores of the factorial images on the original images, i.e. the weight of the linear decomposition, and different pieces of information (such as the eigenvalues) are displayed in a result window.
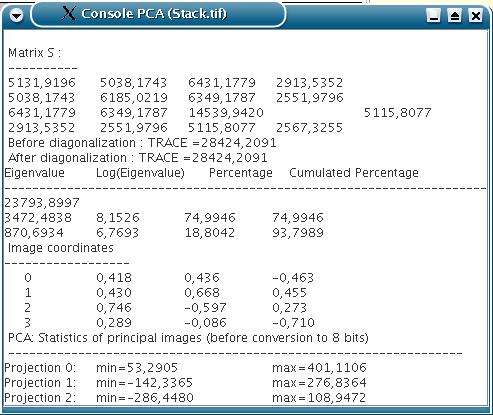
or
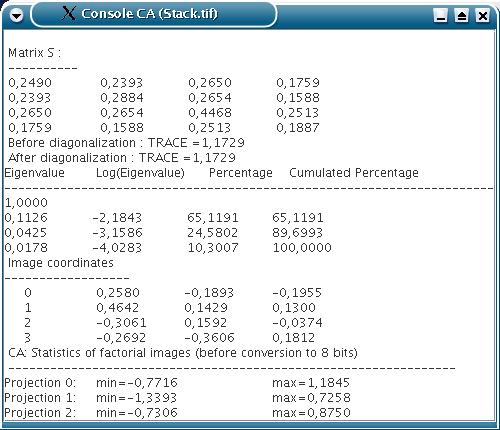
Reduce the dimension of a multivariate data set composed of a series of images.
The series of images may be composed of the different components of a multivariate image (the 3 components of a color image, for instance) or of a set of related images (a time series, for instance).
The series of images is reduced to a smaller series, where the different images of the new series are the principal components (or the factorial images) of the original series.
Method:
Dimensionality reduction is performed according to well-known algorithms (Principal Components Analysis or Correspondence Analysis).
For the moment, only PCA without data centering nor data normalization is implemented. Other variants will be made available soon.
User interface:
Input: The image series to be processed must be within a stack.
An example with four images (four "maps" of an alloy, representing the composition in carbon, chromium, nickel and titanium):




The 4 images are gathered in a stack:

When the plug-in is activated, the user is prompted to select:
a) the method he/she wants to implement (at this moment, only PCA without centering nor mormalization, and CA, are implemented!!!)
b) the number of factorial images he/she wants to get as an output.
The resultant factorial images are put within a stack.
Of course, the stack can be split into factorial images if necessary.
The scores of the factorial images on the original images, i.e. the weight of the linear decomposition, and different pieces of information (such as the eigenvalues) are displayed in a result window.
or