| Authors: |
Leo Goldstien, Daniel Blumenthal, Levi A. Gheber (Levi Gheber Lab)
Department of Biotechnology Engineering, Ben Gurion University of the Negev, Israel |
| Reference: | Universal Approach to FRAP Analysis of Arbitrary Bleaching Patterns |
| How to cite: |
Universal Approach to FRAP Analysis of Arbitrary Bleaching Patterns, Scientific Reports 5, Article number: 11655 (2015)
|
| History: |
2014/09/07: Version 1 .0 - First public release |
| Installation: | Download simFRAP.zip and extract it into the plugins folder inside your ImageJ folder. simFRAP should appear in the plugins drop down list of the ImageJ main window. |
| Source: | Source is available at simFRAP-src.zip or under simFRAP/src in the binary distribution. |
| Description: |
simFRAP allows computation of diffusion coefficients regardless of bleaching geometry used in the FRAP series.
The method is based on fitting a computer-simulated recovery to actual recovery data of a FRAP series.
The algorithm accepts a multiple-frame TIFF file representing the experiment as input and simulates the (pure) diffusion
of the fluorescent probes (2D random walk). Starting with the first post-bleach frame of the actual data, the algorithm iteratively creates a series of simulated images, where each frame corresponds to a single iteration (figure 1).
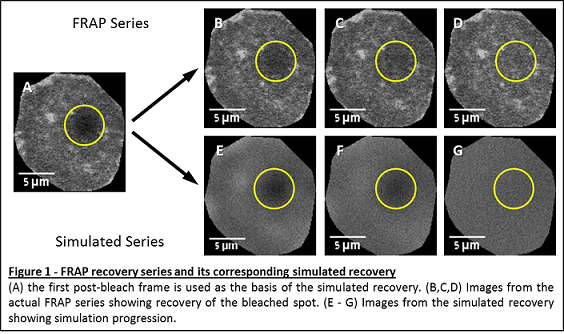
The intensity values are extracted from the user indicated bleached area of the simulated frames, determining the general shape of the recovery curve.
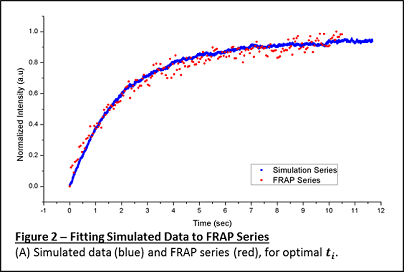
The "time" axis of the simulated data is in arbitrary units (iterations) at this stage. To extract the diffusion coefficient, the simulated recovery curve needs to be fitted to the real recovery curve, by appropriately stretching the "time" axis. An example of simulated data overlapped on real data is shown in Figure 2.
The time between frames in the actual data set is given by the user, allowing the determination of the duration of one iteration in real time units. The diffusion coefficient of the simulated series is then calculated using:
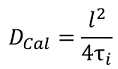 Where l is pixel size (μm) and τi the duration of one iteration (sec). Algorithm requirements:
|
| Sample data: | Download sample data and unzip it in a location where you have write permissions. The zip file contains a FRAP time series acquired in our lab by Daniel Blumenthal of fibroblast cells stained with DiI - a lipid analog. Please refer to the short tutorial contained in the archive for basic usage. |
| Dependencies: | simFRAP makes use of the Apache Commons commons.io.2.4.jar and the JGoodies forms-1.3.0.jar libraries, the source for which is available at their respective sites. |